New research warns that popular deep learning systems trained for cancer pathology may be relying on hidden shortcuts rather than genuine biological signals.
Artificial intelligence (AI) systems designed to predict cancer biology from microscope images may be taking statistical shortcuts rather than detecting genuine biological signals, according to new research from the University of Warwick published in Nature Biomedical Engineering.
The study analyzed more than 8,000 patient samples across breast, colorectal, lung, and endometrial cancers, finding that while AI pathology models often achieved high accuracy rates, they frequently relied on correlations between biomarkers rather than isolating specific biological signals.
“It’s a bit like judging a restaurant’s quality by the queue of people waiting to get in: it’s a useful shortcut, but it’s not a direct measure of what’s happening in the kitchen,” says Dr Fayyaz Minhas, associate professor and principal investigator of the Predictive Systems in Biomedicine Lab in the Department of Computer Science at University of Warwick and lead author of the study, in a release. “Many AI pathology models are doing the same thing, relying on correlations between biomarkers or on obvious tissue features, rather than isolating biomarker-specific signals.”
Models Show Dependence on Correlated Features
The researchers found that instead of directly detecting mutations in cancer-associated genes like BRAF, models often learned that BRAF mutations frequently occur alongside other clinical features such as microsatellite instability (MSI). The systems then used this combination of cues to predict BRAF status rather than learning the actual BRAF signal.
When researchers assessed AI model performance within specific patient subgroups—such as only high-grade breast cancers or only MSI-positive tumors—accuracy fell substantially. This revealed the models’ dependence on shortcut signals that disappear when confounding factors are controlled.
“We’ve found that predicting a BRAF mutation by looking at correlated features like MSI is often like predicting rain by looking at umbrellas—it works, but it doesn’t mean you understand meteorology,” says Kim Branson, senior vice president global head of artificial intelligence and machine learning at GSK and co-author, in a release.
Limited Performance Advantage Over Clinical Data
For certain prediction tasks, the performance advantage of deep learning over human-derived clinical information was modest. AI systems achieved accuracy scores of just over 80% when predicting biomarkers, compared with around 75% using tumor grade alone—a measure already assessed by pathologists.
“Crucially, if a model cannot demonstrate information gain above a simple pathologist-assigned grade, we haven’t advanced the field; we’ve just automated a shortcut,” says Branson in a release. “The roadmap for the next generation of pathology AI isn’t necessarily bigger models; it’s stricter evaluation protocols that force algorithms to stop cheating and learn the hard biology.”
The findings raise concerns about deploying current AI pathology tools in routine patient care without stronger evaluation standards. The researchers call for approaches that explicitly model biological relationships and causal structure, along with subgroup testing and comparison against clinical baselines.
“This research is not a condemnation of AI in pathology. It is a wake-up call,” says Minhas in a release. “Current models may perform well in controlled settings but rely on statistical shortcuts rather than genuine biological understanding. Until more robust evaluation standards are in place, these tools should not be seen as replacements for molecular testing.”
The researchers note that machine learning methods can still prove valuable for research, drug development screening, and clinical decision support, but emphasize the need for bias-aware evaluation rather than relying solely on headline accuracy metrics.
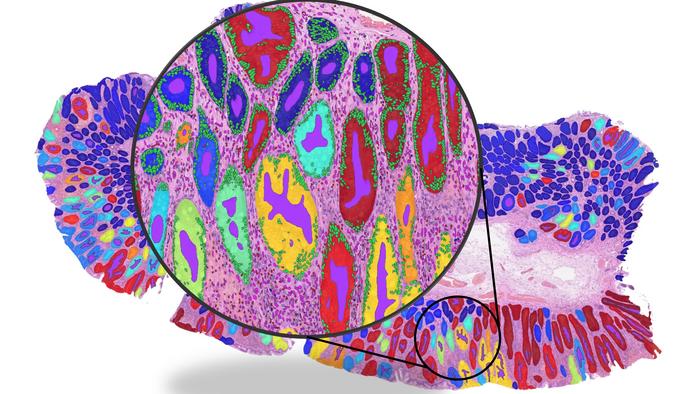
Photo caption: Whole slide image illustrating the detection of key histological structures such as glands and cells.
Photo credit: Dr Fayyaz Minhas / University of Warwick
We Recommend for You: