Advanced methods of molecular analysis promise breakthroughs in diagnosis and guided therapy
By Christoph Menzel and Michael Kazinski
In modern medical practice, the use of tissue biopsies is a well-established method for diagnosing cancer and other diseases. However, such procedures come with certain technical limitations that can make it impossible to obtain enough sample material from patients. In addition, the method is ill suited for regular monitoring to gauge treatment response.
In recent years, researchers have begun to make use of advanced molecular technologies to develop techniques for performing “liquid biopsies,” using markers found in blood or other body fluids to overcome the limitations of tissue biopsies. By enabling clinicians to detect and analyze molecular biomarkers found in body fluids, this emerging method holds significant potential for profiling the tumors where the biomarkers originated, and thereby improving the diagnosis and treatment of cancer and other diseases.
THE GOLD STANDARD, FOR NOW

Figure 1. The use of tissue biopsies for cancer diagnosis is widespread, yet the practice is burdensome, expensive, and carries risks of infection or other health complications. Photo courtesy Qiagen and Andreas Fechner.
The use of tissue biopsies for cancer diagnosis is common and widespread. In the United States, for example, it is estimated that 1.6 million breast biopsies and 1 million prostate biopsies are performed each year.1 Regardless of whether such procedures are performed as highly invasive surgery, as endoscopic examinations, or as needle punctures, each tissue biopsy represents an invasion of the patient’s body. And all such invasive approaches carry the risk of infection or other complications for the patient (see Figure 1).
Even when invasive procedures are used, it can be challenging to obtain sufficient tissue to perform all the diagnostic procedures that may be required—including molecular biomarker testing to help guide treatment decisions for targeted therapies. In cases where enough tissue is available, a traditional biopsy might still not reveal all of the information necessary to diagnose a patient’s condition, in part because of the inherent heterogeneity of tumor tissue. Such limitations are particularly true for advanced metastatic stages of cancer.
On top of the challenges involved in obtaining adequate patient specimens, tissue biopsies are also costly procedures. The high costs of performing tissue biopsies are partly due to the invasive nature of the sample collection process. But the costs are also influenced by the lengthy workflows necessary to process a tissue biopsy sample—typically by fixation in formalin and embedding in paraffin—as a prerequisite for downstream analysis.
Analyses that use tissue biopsy specimens to create a detailed molecular characterization of the tumor are becoming vital for state-of-the-art disease management. Today, numerous genomic alterations are recognized as markers for the diagnosis, prognosis, and monitoring of cancer, or as predictive markers for the patient’s response to a targeted therapy. To guide treatment decisions, for example, clinicians routinely use a variety of molecular methods to test for mutations of the KRAS gene in colorectal cancer, the BRAF gene in melanoma, and EGFR mutations and ALK fusions in lung cancer.
In addition, FDA has already approved more than 20 companion diagnostic tests, and many more are expected in the coming years. For now, the majority of such tests are based on formalin-fixed, paraffin-embedded tissue biopsies—with all the disadvantages associated with such sample materials.
IMPROVING DIAGNOSIS AND MONITORING
Liquid biopsies represent a new, minimally invasive technique that enables the detection and analysis of molecular biomarkers in blood and other body fluids. Researchers and test developers believe that the new technique holds great promise for overcoming the shortcomings of tissue biopsies.
Figure 2. Minimally invasive liquid biopsies based upon blood, urine, or other body fluids hold promise for improving diagnosis and patient care for many conditions. Photo courtesy Qiagen and Andreas Fechner.
In general, liquid biopsy technologies enable researchers and clinicians to employ a quick and comparatively straightforward process that usually includes a simple blood draw, extraction of nucleic acids from the blood plasma, and amplification of the molecular targets, to enable analysis of defined biomarkers (see Figure 2).
A major advantage of liquid biopsies is that physicians can use them repeatedly to monitor changes in tumor genetics over time—a process that is difficult or impossible to carry out using tissue biopsies. Even the newest and most effective personalized cancer therapies frequently result in drug resistance and relapse of the tumor. A reassessment of the tumor’s molecular profile is highly beneficial in such cases, but obtaining a new tissue sample is often impossible. The liquid biopsy approach overcomes this difficulty, and—in combination with the newest molecular characterization methods, such as next-generation sequencing—holds great promise to improve understanding of disease progression.
Although blood or other body liquids constitute the sample materials needed to perform liquid biopsies, the detailed source of the genetic information within the sample may be different. At present, researchers recognize three main sources for the DNA and RNA extracted from liquid biopsies: circulating tumor cells, free circulating DNA, and exosomes.
Tumor cells originating from a primary tumor were first observed in circulating blood more than a hundred years ago. Nevertheless, such circulating tumor cells (CTCs) are usually rare. Even in metastatic stages of cancer progression, only a few cells are typically found in several milliliters of plasma. Consequently, special methods for enrichment of the tumor cells—or depletion of non-tumor cells—are needed prior to molecular characterization.
A more recent approach to liquid biopsy is based on circulating cell-free nucleic acids, which are believed to originate mainly from dead cells. Circulating free tumor DNA (cfDNA or ctDNA) has been shown to contain cancer-related mutations. Researchers analyze ctDNA together with plasma DNA from normal cells, which is always present in the circulation. Once again, a highly effective enrichment of the circulating free DNA is required.
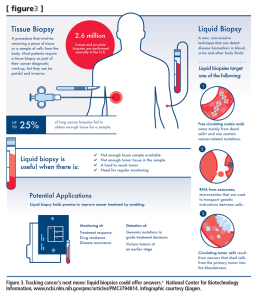
Figure 3. Tracking cancer’s next move: liquid biopsies could offer answers. a National Center for Biotechnology Information, www.ncbi.nlm.nih.gov/pmc/articles/PMC3794814. Infographic courtesy Qiagen.
One of the latest approaches to liquid biopsies focuses on the detection of nucleic acids from exosomes. Exosomes are microvesicles shed by cells in the body as part of a biological communication system. Each exosome can carry a tiny cargo of genetic instructions in the form of DNA and RNA molecules through the blood, urine, or other fluids. Exosome detection may prove very useful in diagnosing and monitoring diseases, including brain cancer, where the blood-brain barrier inhibits the emission of cfDNA into the bloodstream (see Figure 3).
CHALLENGES FOR LIQUID BIOPSIES
Conceptually, liquid biopsies hold great promise for overcoming many of the limitations associated with tissue biopsies. In practice, however, the use of liquid biopsies may still encounter some challenges related to the comparatively low concentration of molecular biomarkers in body fluids. Many of these challenges can be effectively addressed by adjusting the laboratory workflow, as described below.
A number of technologies and approaches are available for isolating CTCs from a patient’s blood sample. Because the concentration of CTCs in such samples is usually very low, however, the amount of DNA or RNA that such capture technologies can recover by themselves is severely limited. In order to perform downstream analyses, including whole-genome sequencing, it is necessary to perform an unbiased amplification of the sample’s genomic content.
The low concentration of target DNA in a plasma sample is also a major challenge when working with cfDNA. To ensure the highest sensitivity of downstream analyses, it is critical to achieve the highest possible recovery of cfDNA, since this will also increase the number of true positives captured. To increase the amount of cfDNA available for analysis, it is necessary to draw a larger volume of blood from the patient.
A larger blood draw will significantly improve the recovery of DNA from the sample. In the case of early-stage tumors, for example, a plasma sample of approximately 2 ml may yield a sensitivity of only 50%. Increasing the size of the sample to 4 ml or greater (to as much as 10 ml), can dramatically improve the sensitivity of downstream analyses to a range of 80% to 90%.
Proper handling and storage of the sample is also critical. If the sample is not sufficiently stabilized, for example, background DNA can be released from white blood cells. Reducing such background DNA helps to ensure that the appropriate sensitivity can be obtained. Special storage tubes are available to help mitigate the effects of background DNA. And blocking oligonucleotides can also be used to ensure that the background DNA isn’t amplified or sequenced, thereby minimizing the effects of background DNA.
In serum or plasma samples, circulating free nucleic acids are usually present in the form of short fragments. In the case of DNA, for example, the fragments are typically less than 1000 bp in length. To ensure the appropriate yield and purity of recovered DNA, a dedicated preparation process should be used on samples found to contain fragmented DNA. Yield and purity may also be maximized through process standardization and testing automation. The QiaAmp DSP circulating nucleic acid kit, for example, offers a standardized means of purifying such circulating nucleic acids from human plasma, serum, or other cell-free body fluids.
Exosomes are yet another source of circulating nucleic acids that may yield valuable diagnostic information. Exosomes offer great advantages as source materials, as they preserve DNAs, RNAs, and even proteins from the degrading extracellular environment, helping to make it possible to understand disease mechanisms by analyzing their transcriptomic profile. However, these microvesicles are excreted from cells, and are therefore smaller than cells. Capturing just the exosomes and not any other cellular material, while also maintaining the integrity of the nucleic acids, poses a significant challenge.
APPLICATIONS IN CLINICAL RESEARCH AND HEALTHCARE
Despite the practical challenges involved, liquid biopsies hold great promise for improving diagnosis, treatment, and outcomes for patients with a variety of diseases. Initial applications of the technique are being sought especially for several types of cancers. Lung cancer is a prime example of how the potential benefits of liquid biopsies can apply for healthcare professionals and patients.
In advanced lung cancer, performing a tissue biopsy is often prohibited because the patient’s condition is so poor that the procedure is too risky. It may also be impossible to perform a tissue biopsy because the tumor or metastases is sitting in a part of the lung that is inaccessible even with CT-guided fine needle aspiration. And even if a core needle biopsy or fine needle aspiration can be performed successfully, after various pathological and histological examinations the remaining tumor material may not be sufficient for performing molecular companion diagnostic testing to detect EGFR mutations.
As a result of such obstacles, for many lung cancer patients there are either no tissue samples available at all, or the low quantity or poor quality of the existing samples makes them unusable. It is estimated that a tissue biopsy is not evaluable in up to 25% of cases of advanced non-small cell lung cancer (NSCLC). Without the analyses that might be based on such samples, treatment options for individual patients may be extremely limited.2
With the emergence of liquid biopsies, these scenarios are just beginning to change. Early clinical studies focused on NSCLC patients have shown that predictive testing can be achieved by means of liquid biopsies performed on blood samples, as a surrogate for determining the patient’s EGFR status from tissue samples. In a recent study of the tyrosine kinase inhibitor gefitinib, for instance, it was found that EGFR mutation status could be accurately assessed using circulating-free tumor DNA, with high specificity and sensitivity and a high concordance to the tissue result.2 Similar studies are under way for targeted therapies in colorectal and skin cancer.
A LOOK INTO THE FUTURE
Ongoing research is highlighting the great promise of liquid biopsy technologies across a number of fields. To expand the range of clinical applications, a growing number of joint initiatives involving pharmaceutical and diagnostic companies are aimed at developing novel companion diagnostics based on liquid biopsies.
Many researchers are now seeking new applications for liquid biopsy technologies. The use of liquid biopsies appears to be highly attractive for improving disease monitoring and eventually even as a tool for cancer screening. Especially in the realm of cancer diagnosis and therapy, the use of liquid biopsies to achieve highly specific diagnoses and to guide corresponding personalized therapies closely holds the potential of significantly improving outcomes for many cancer patients.
Christoph Menzel, Dr.rer.nat., is director of global product management for molecular diagnostics and personalized healthcare/oncology, and Michael Kazinski is senior director for molecular preanalytic sample technologies, at Qiagen. For further information, contact CLP chief editor Steve Halasey via [email protected].
REFERENCES
- Grady D. Study of breast biopsies finds surgery used too extensively. New York Times, February 18, 2011. Available online: www.nytimes.com/2011/02/19/health/19cancer.html. Accessed April 26, 2015.
- Douillard JY, Ostoros G, Cobo M, et al. Gefitinib treatment in EGFR mutated caucasian NSCLC: circulating-free tumor DNA as a surrogate for determination of EGFR status. J Thorac Oncol. 2014;9(9):1345–1353; doi: 10.1097/JTO.0000000000000263.








Cell-free DNA based liquid biopsy is the ultimate solution to overcome all limitations of tissue biopsy. However, current version of liquid biopsy requires 10-20 mL of blood due to unavoidable sample loss in cfDNA recovery. Losing a lot of starting materials mean you will compromise the accuracy of the end results in a clinical setting. CirculoGene has come up with a unprecedented finger-stick liquid biopsy enables physicians to work with a sample volume as little as 20 uL!!! It’s crazy the pace that technology advancing.