A team of researchers from the Italian Institute of Technology has designed in silico “molecular probes” able to track the progress of a protein that misbehaves in different neurodegenerative diseases, such as Amyotrophic Lateral Sclerosis (ALS) and Fronto-Temporal Dementia (FTD).
The probes can be used to study the behavior of the target protein in cell and were tested in collaboration with Sapienza University of Rome, Centre for Genomic Regulation n Barcellona, University of Edinburgh and Kings College London. The research study has been published in Nature Communications.
Created by the “RNA Systems Biology” group at IIT in Genoa, the probes consist of computer-designed RNA molecules that bind to a neurodegeneration-associated protein named TDP-43. This protein is present in numerous cases of Amyotrophic Lateral Sclerosis (ALS) and Fronto-Temporal Dementia (FTD), where it aggregates creating insoluble protein blobs in neural cells, altering their metabolism and function.
The research team was inspired by the protein’s natural interactions with RNA molecules to design molecular probes, which are called “aptamers,” literally molecules made to fit one single target. Their main goal was to obtain a novel approach for tracking the aggregation of neurodegeneration-associated proteins at the very first steps of the process.
“Using our own algorithms, we designed RNA aptamers specific for TDP-43 and used them together with advanced microscopy techniques to follow the protein transition towards its aggregated forms” says Gian Gaetano Tartaglia, principal investigator of the RNA System Biology Lab. “We can identify TDP-43 aggregates as small as 10 nanometers which, to our knowledge, is the best resolution achieved so far when visualising protein aggregates”.
These aptamers could be used to study, at the molecular level, the phenomenon of abnormal protein aggregation typical of several neurodegenerative diseases and would, therefore, pave the way for the development of early diagnosis tools for these disorders.
“We showed that the RNA aptamers can also be used to track TDP-43 in live cells and in real time, detecting all forms of the protein, from the physiological soluble one to the insoluble state, passing by aggregates of intermediate sizes undetectable by standard approaches,” says Elsa Zacco, lead researcher on the project.
The study was carried out by IIT researchers Elsa Zacco, Alexandros Armaos and Gian Gaetano Tartaglia (also at Sapienza University of Rome), with the participation of the groups led by Mathew Horrocks at University of Edinburgh and Annalisa Pastore at King’s College London.
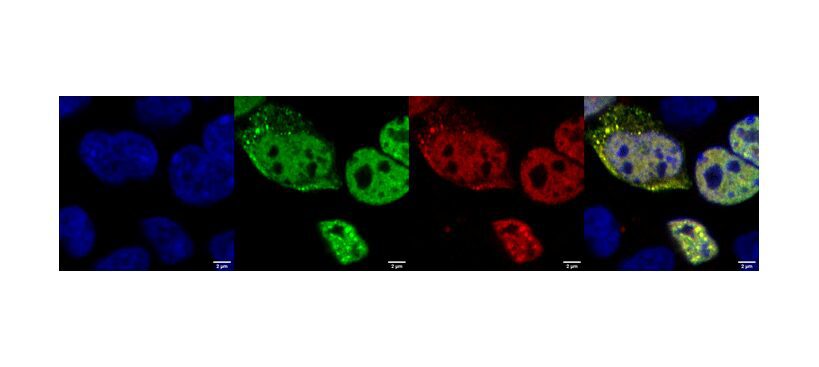
Featured image: Zoom-in on single cultured mammalian cells in which TDP-43 has been induced to aggregate. In this system, the cells produce TDP-43 fused to a green fluoresce molecule, to be able to detect whether the protein forms insoluble granules (green fluorescent dots). The RNA probe is labelled with a red fluorescent tag. The yellow color, given by the overlap between the green of TDP-43 and the red of the RNA probe, signifies that the probe can search and find its protein target in live cells, suggesting that it could be used as a detection tool to track the progress of TDP-43 aggregation in disease. Blue: nuclei; green: TDP-43; red: RNA probe; yellow: TDP-43+RNA probe. Photo: Italian Institute of Technology