New data show a combined clinicopathologic and gene expression profile test outperforms standard risk assessment tools in identifying sentinel lymph node metastasis risk in early-stage melanoma.
New data published in the Journal of the American Academy of Dermatology demonstrate that a combined clinicopathologic and gene expression profile (CP-GEP) test can identify high-risk sentinel lymph node (SLN) metastasis in T1a melanoma patients—a population traditionally considered low risk—with greater specificity than conventional assessment methods.
The analysis, led by Dr Wesley Yu et al and published in October 2025, found that the Merlin CP-GEP test identified 18% of T1a melanoma patients as high risk, with SLN positivity rates exceeding commonly accepted biopsy thresholds. Compared to the MIA nomogram using a 5% threshold, CP-GEP demonstrated superior specificity (72.0% vs 23.7%) while maintaining a negative predictive value greater than 98%, according to a release from SkylineDx, the test’s developer.
“CP-GEP identified a subset of T1a patients with SLN positivity above guideline thresholds (>10%), enabling actionable risk stratification,” the authors concluded, in a release.
Biology-Driven Test Design
Merlin CP-GEP integrates eight biologically selected genes associated with metastatic pathways, originally identified and developed in collaboration with investigators at Mayo Clinic. The assay combines clinicopathologic variables with gene expression profiling into a single integrated algorithm, offering binary stratification of patients into high- or low-risk categories for metastasis.
The test is the only commercially available gene expression profile assay that integrates clinicopathologic factors with gene expression profiling, according to SkylineDx. Findings from the prospective MERLIN_001 trial, published in JAMA Surgery in October 2025, similarly identified a subset of T1a patients with elevated risk, corroborating the biologic heterogeneity observed in the newly published analysis.
NCCN Guideline Recognition
The National Comprehensive Cancer Network has updated its Clinical Practice Guidelines in Oncology for Cutaneous Melanoma (Version 1.2026) to recognize the CP-GEP test as a predictive genomic test that may be used to support metastatic risk assessment in T1b and T2a melanoma patients. The guidelines differentiate Merlin CP-GEP from other gene expression profile assays, stating that alternative tests are not recommended for sentinel lymph node biopsy (SLNB) risk prediction outside of clinical trials, according to SkylineDx.
As demonstrated in the MERLIN_001 trial, a patient identified as high risk by CP-GEP may have approximately a threefold increased likelihood of SLN positivity, per the company.
Competing Assay Shows No High-Risk Classifications in T1a Patients
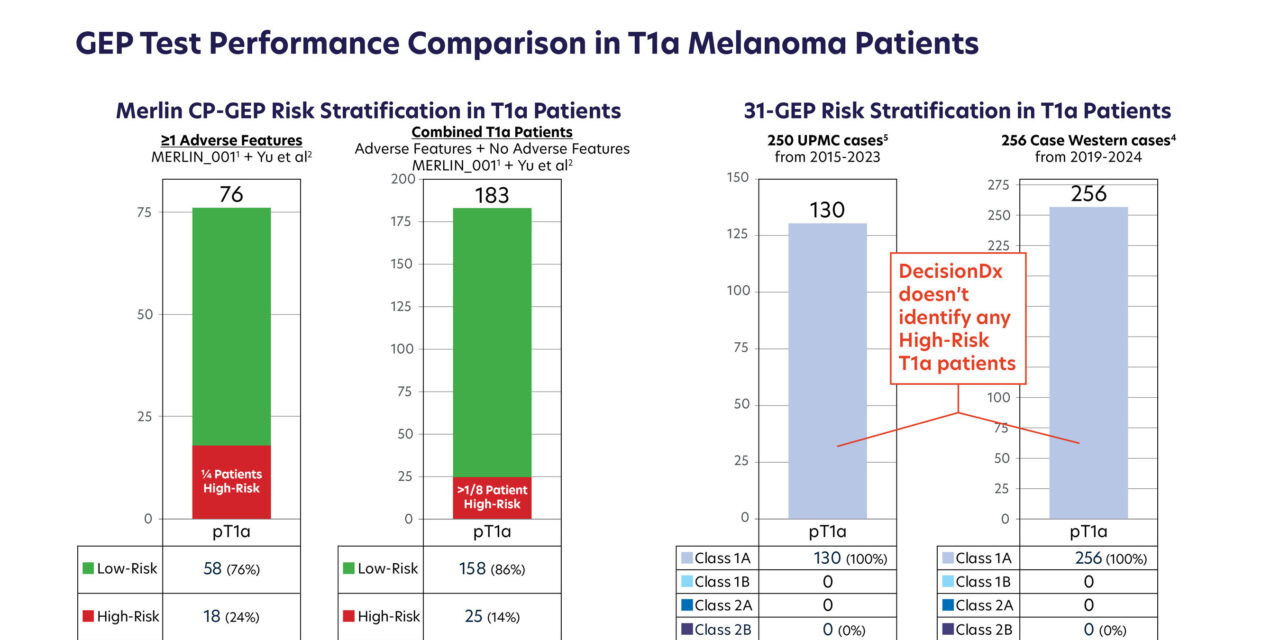
Two independent publications evaluating a separate 31-gene expression profile assay in exclusively T1a melanoma populations reported no high-risk classifications across a combined cohort of 506 patients—250 from the University of Pittsburgh Medical Center (2015–2023) and 256 from Case Western Reserve University (2019–2024).
In both cohorts, 100% of T1a tumors were classified as the lowest-risk category.
“In our cohort, 100% of 256 pT1a tumors were GEP-31-Class 1A,” the authors of one study reported, in a release. “Our findings do not support 31-GEP testing in this group.”
The current CMS/Medicare reimbursement rate for the 31-gene expression profile test exceeds $7,000 per test, according to SkylineDx, underscoring the importance of demonstrating clear clinical value when no high-risk classifications were identified.
Addressing Mortality in Early-Stage Disease
More than 50% of melanoma deaths in the US arise from early-stage disease, according to data from the SEER registry. A substantial proportion of T1a patients with adverse biologic features may not currently undergo SLNB under standard practice patterns, according to SkylineDx.
By identifying biologically aggressive tumors that exceed guideline SLNB thresholds, the CP-GEP test aims to give clinicians additional clarity when deciding whether to recommend SLNB for patients who might otherwise be considered low risk under traditional clinicopathologic criteria alone.
Photo caption: GEP test performance comparison in T1a melanoma patients
Photo credit: SkylineDx
Related Reading:
Studies Find Merlin CP-GEP Test Improved Melanoma Risk Stratification
Large Melanoma Trial Validates Gene Expression Test’s Predictive Accuracy
AI Model Achieves 94.5% Accuracy in Melanoma Detection Using Combined Data Sources