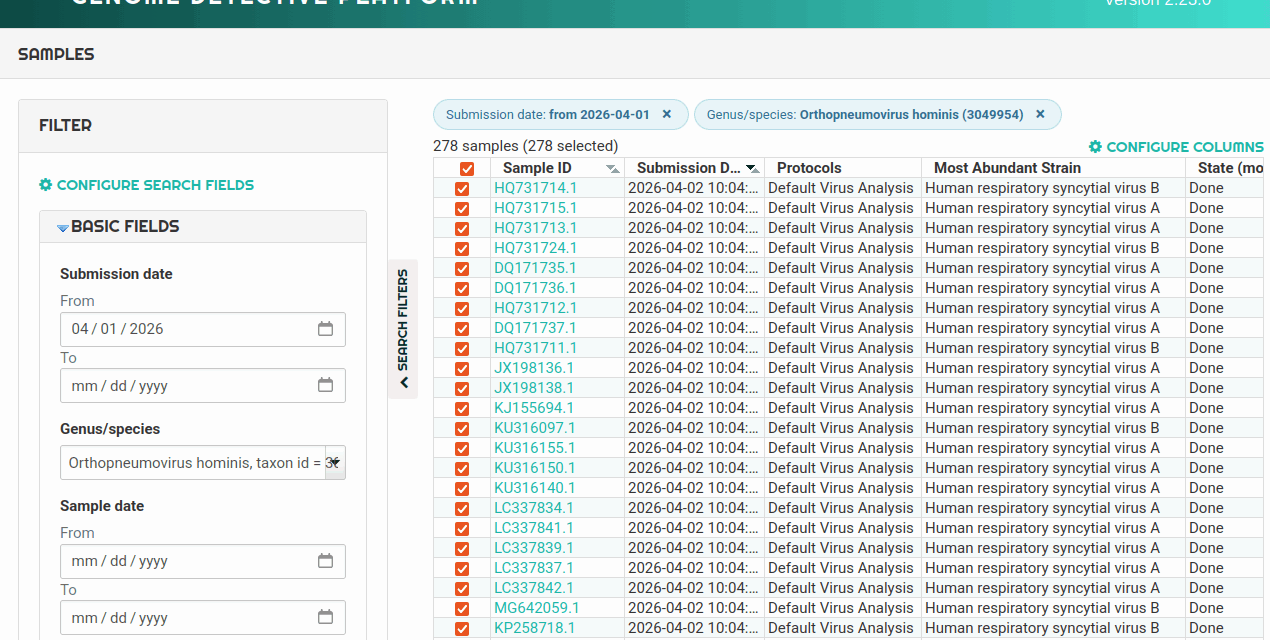
The cloud-based system aims to address the bottleneck of translating raw genomic data into actionable information for clinical laboratories.
Genome Detective will present its automated pathogen genomics platform at the European Society of Clinical Microbiology and Infectious Diseases (ESCMID) Global 2026 in Munich, April 17-21. The system is designed to help clinical and public health laboratories translate raw next-generation sequencing (NGS) data into standardized and actionable information.
The cloud-based bioinformatics environment automates the identification and characterization of viral and bacterial pathogens from NGS datasets. The platform integrates quality control, assembly, alignment, classification, typing, and variant detection into a single workflow. Key technical features include automated taxonomic classification, optimized assembly pipelines, variant detection, and antimicrobial resistance gene detection.
“Genome Detective combines ease of use with high analytical quality, delivering reliable results without requiring bioinformatics expertise. Its transparent data and accuracy give us confidence for consistent, trustworthy decision-making in routine microbiology,” says Stefan A Boers, medical molecular microbiologist, in a release.
Reducing Bioinformatics Dependency
By automating computational steps and providing standardized outputs, the platform aims to reduce operator variability and enhance reproducibility, which are key factors for accreditation and quality assurance frameworks. The system generates results within minutes to hours, depending on the complexity of the dataset, and presents them in structured formats.
The scalability of the platform allows both smaller laboratories and national reference centers to implement genomic analysis without building custom bioinformatics infrastructure. It is currently deployed for applications including rapid pathogen identification from clinical samples, genomic outbreak investigation, viral variant surveillance, and metagenomic pathogen discovery.
“Reflecting on the COVID-19 pandemic, the critical need for fast and accurate bioinformatics software is clear. We could not have achieved what we did without the support of Genome Detective,” says Tulio de Oliveira, director CERI and Krisp, in a release.
Scientific Presentations at ESCMID Global
During the conference, the Genome Detective team will demonstrate current workflows and discuss integration into diagnostic pipelines and public health reporting structures. The platform is currently a research-use-only (RUO) tool for pathogen genomics.
The company will also contribute to the scientific program with a presentation on April 20 regarding the acceleration of pathogen identification from NGS data. Additionally, two posters will be presented:
P0367: Evaluation of the platform as a tool for viral epidemiological surveillance.
P3264: A web-based pipeline for automated fungal identification via targeted sequencing.
“The reliability and continuous development of Genome Detective give us strong confidence in our results. It is now fully integrated into our workflow and significantly strengthens our capacity to respond to re-emerging viral agents,” says Cintia Fabbri, researcher at the Instituto Nacional de Enfermedades, in a release.
Photo caption: Genome Detective
Photo credit: Genome Detective
Related Reading: