The Fungal Reference Laboratory at the University of Alabama at Birmingham (UAB) developed its own COVID-19 diagnostic test and used pooled testing to enable a safe return to college campuses throughout the state of Alabama in summer 2020. Now, the lab is using what it learned about the virus to identify its variants:
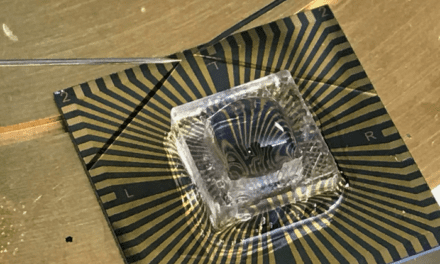
This is relatively simple in a facility such as UAB’s Fungal Reference Laboratory, directed by Sixto Leal, M.D., Ph.D., assistant professor in the Department of Pathology. UAB has three Thermo Fisher Genexus systems capable of sequencing up to several hundred viral genomes per week, Leal said. “This sequencing can detect any variant out there.”
Once they have a sequence, Leal and his team compare it with the SARS-CoV-2 reference genome and to lists of known variants.
At UAB, Leal explains, patient samples are sequenced if the patient:
- has a severe or atypical presentation of COVID-19 (or MIS-C in the case of pediatric patients); or
- has a positive COVID-19 test after full vaccination; or
- exhibits an S dropout on the Thermo Fisher TaqPATH PCR test (more on that below); or
- has epidemiologic links suggestive of possible exposure to the South African, Brazilian, New York or California variants.
When a variant is identified, Leal’s team immediately notifies the Alabama Department of Public Health, which begins intensive contact tracing. “If you identify a patient or cluster of patients with a particular variant, you may be able to stop it from spreading throughout the state,” Leal said.
Read from the University of Alabama at Birmingham.
Featured image: UAB’s Fungal Reference Laboratory uses three Thermo FIsher Ion Torrent Genexus NGS systems like this to identify COVID-19 variants.