Scientists at the Scripps Research Institute (TSRI), La Jolla, Calif, have analyzed a key protein from two influenza strains that recently began causing sporadic infections among people in China and Taiwan.
The analyses suggest that the flu viruses, variants of subtypes H10N8 and H6N1, have not acquired changes that would allow them to infect people easily and cause a much-feared pandemic.
Yet the studies also highlight the versatility that bird flu viruses apparently have in attaching to host cells.
“These bird flu viruses seem able to bind to receptors on host cells in different ways, and thus can probably mutate in different ways to jump to humans—so we shouldn’t be complacent about our ability to predict the viral changes required to get a pandemic,” says Ian A. Wilson, PhD, Hansen professor of structural biology and chair of TSRI’s department of integrative structural and computational biology.
The new research, from Wilson’s laboratory and the laboratory of James C. Paulson, PhD, acting president and CEO of TSRI, was reported in two papers in the journal Cell Host & Microbe, on March 11, 2015.
WEAK BINDING
One study focused on an H10N8 bird flu virus that infected and killed a 73-year-old Chinese woman from Jiangxi province in late 2013, and has reportedly caused at least two other deaths in that province since then.
Using the genetic sequence of the virus obtained by Chinese scientists, the TSRI team produced copies in cell culture of its hemagglutinin (HA) protein, which flu viruses use to infect host cells.
The influenza HA protein mediates infection by first binding to the tips of sugar-like molecular chains that extend from certain cells—including epithelial cells that line the airway and gut. Paulson’s laboratory tested the ability of the human-infecting H10N8 HA protein to bind to lab-grown versions of these ‘flu virus receptors’ and found that it bound well to the receptors found on bird cells, but negligibly to those found on human cells.
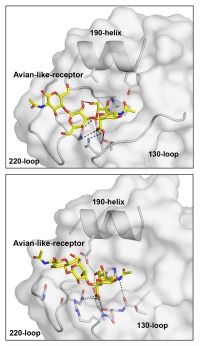
A new analysis suggests that two deadly flu viruses, variants of subtypes H10N8 and H6N1, have not acquired changes that would allow them to infect people easily. Image courtesy the Scripps Research Institute.
Meanwhile Wilson’s laboratory used x-ray crystallography to determine the atomic-scale structure of the H10N8 HA protein—on its own, while bound to an avian-like receptor, and while bound weakly to a human-like receptor. These studies revealed precise structural details of how the human-infecting H10N8 HA binds to avian receptors and why it cannot bind well to human receptors.
“This H10N8 HA protein hasn’t acquired the structural changes that would allow the virus to be transmitted efficiently from human to human,” says Heng Zhang, a research associate in the Wilson laboratory who was a co-first author of the H10N8 paper, together with Robert de Vries and Netanel Tzarum.
Intriguingly, although the viral gene sequence data indicated that the sporadically human-infecting H10N8 HA had mutated slightly away from ordinary bird-infecting H10N8 HA, the data suggested that none of these mutations had materially improved the HA protein’s ability to bind to human receptors. And when Paulson’s team introduced new mutations into the HA mutations that have caused other bird flu subtypes to switch their preference from bird receptors to human receptors, the altered HA still failed to bind well.
The other study, of the HA protein from an H6N1 flu virus that caused nonfatal illness in a 20-year-old Taiwanese woman in June 2013, came to essentially the same conclusions: the viral HA had changed somewhat compared to the usual HA from bird-isolated H6N1, but still remained adapted for binding to avian, not human, receptors.
DIFFICULT TO PREDICT
How did such bird viruses end up causing infections of people? “We suspect that sporadic cases of human infection by a bird flu virus can occur, even without a change in the receptor specificity, if the dose of the viral exposure is high enough and/or it gets deep into the lungs, where there are some flu-virus receptors like those found in birds,” says Tzarum, a research associate in the Wilson laboratory who was first author of the H6N1 paper.
Flu viruses with these HAs thus remain essentially bird viruses, with limited ability to infect humans. Yet further mutations that would enable a switch in preference to human receptors—and a potential global pandemic—are still possible.
The new TSRI analyses also show that, at the atomic scale, these new bird flu HAs bind to host-cell receptors in ways not observed in studies of other bird flu viruses—implying that the mutations required for the switch to human receptors may be different for different strains and inherently hard to know in advance.
“There appear to be no general rules for this switch among bird flu viruses,” says Tzarum.
Determining whether a bird flu strain has truly jumped the species barrier will therefore continue to require detailed receptor-binding and structural studies like these, Wilson says.
The other co-authors on both papers were Robert P. de Vries, Xueyong Zhu, Wenli Yu, and Ryan McBride, all of TSRI at the time of the research. The studies were funded in part by the National Institutes of Health.
For more information, visit the Scripps Research Institute.
REFERENCES
Tzarum N, De Vries RP, Zhu X, et al. Structure and receptor binding of the hemagglutinin from a human H6N1 influenza virus. Cell Host & Microbe. 2015; 17(3):369–376; doi: http://dx.doi.org/10.1016/j.chom.2015.02.005.
Zhang H, de Vries RP, Tzarum N, et al. A human-infecting H10N8 influenza virus retains a strong preference for avian-type receptors. Cell Host & Microbe. 2015; 17(3): 377–384; doi: http://dx.doi.org/10.1016/j.chom.2015.02.006.








