Researchers integrate genomic, transcriptomic, and proteomic data with metabolic modeling to create machine learning classifier for lung and pancreatic cancers.
Researchers from Yeditepe University in Turkey have developed a systems biology workflow that combines multiple types of molecular data with metabolic modeling to improve cancer diagnosis and subtyping accuracy.
The approach, published in Quantitative Biology, integrates patient transcriptomic data into human genome-scale metabolic models to generate personalized metabolic flux distributions. These flux profiles are then combined with genomic, transcriptomic, and proteomic data to train a multi-omic machine learning classifier.
“Traditional diagnostic methods like biopsies are invasive and carry risks, while non-invasive approaches often struggle to identify reliable biomarkers due to tumor heterogeneity and the complexity of biological data,” according to the research team.
High Accuracy in Lung Cancer Detection
The integrated approach was primarily tested on lung cancer data, achieving high accuracy in distinguishing cancer from normal tissue, differentiating subtypes such as adenocarcinoma versus squamous cell carcinoma, and identifying early-stage disease.
The classifier identified key biomarker features across all molecular layers, including metabolic pathways like lipid and amino acid metabolism that had not been previously highlighted in cancer diagnosis.
When applied to pancreatic cancer data, which typically suffers from limited sample availability, the workflow demonstrated strong performance, proving its robustness across different cancer types.
Metabolic Modeling Integration
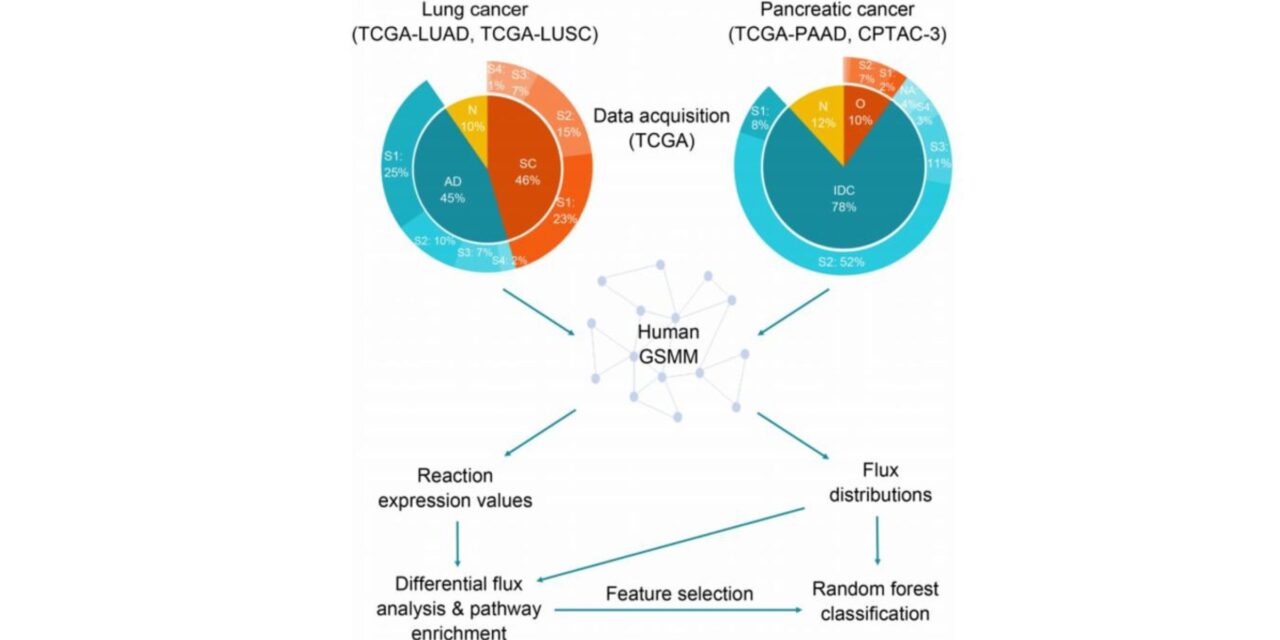
The workflow acquires multi-omic data from sources like The Cancer Genome Atlas and processes transcriptomic data through genome-scale metabolic models to compute patient-specific flux distributions. These flux profiles, representing the functional metabolic state of cells, are combined with genomic and proteomic data to build the multi-omic classifier.
The approach addresses limitations of isolated data analysis by capturing the dynamic metabolic state of tumors, which has been technically challenging in previous methods.
Photo caption: The integrated systems biology workflow for cancer subtyping and early diagnosis. The pipeline combines multi-omic data with metabolic modeling to generate patient-specific flux distributions, which are used to train a high-performance classifier and identify critical biomarker pathways. (Concept based on Tanil & Nikerel, Quantitative Biology, 2025).
Photo credit: Higher Education Press